Laboratory of Molecular Biology
PROFILE
Our research is focused on the analysis of molecular mechanisms responsible for DNA replication in bacterial cells. We mainly investigate the mechanisms responsible for the initiation and regulation of DNA replication of bacterial plasmids an antibiotic resistance factors in bacterial cells. Model used in our investigations is RK2 plasmid that belongs to the group of broad-host-range replicons that can be stably maintained and transferred between cells of various bacterial species, including plant, animal and human pathogens. The research utilizing such a unique system, allowing for the analysis of cellular mechanisms in various organisms, makes the significant contribution to-wards the general understanding of the fundamental cellular processes im-portant for future development of new antimicrobial therapeutic strategies.
In our investigations we focus on (i) multi-molecule complexes formed on DNA by replication initiation proteins (Rep), their structural bases, functions and dynamics, (ii) analysis of the mechanism of protease activity on substrates bound to DNA and effects of proteases on DNA replication regulation and stably maintenance of plasmids, and (iii) analysis of the function of plasmid encoded post segregational killing systems in the context of its components’ interactions with DNA and their stability.
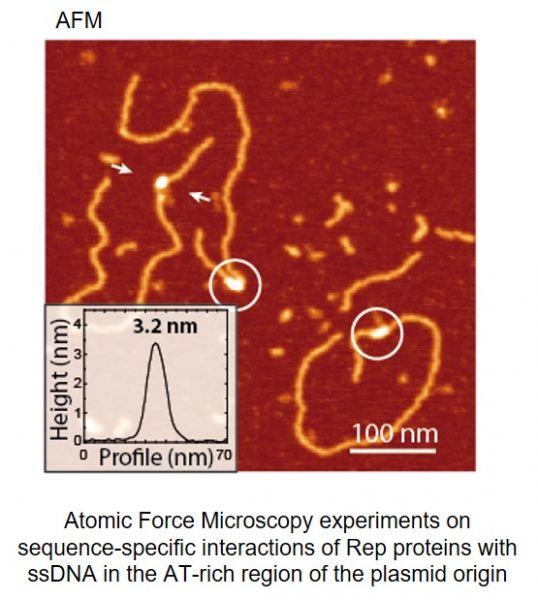
Current understanding of structure and functions of multi-protein complex-es formed on DNA is still limited. Especially challenging is the analysis of in-trinsically disordered proteins and their complexes with single-stranded DNA (ssDNA). After discovery that Rep proteins bound both double-stranded DNA (dsDNA) and similar, as chromosomal DnaA, they also interact with ssDNA (Wegrzyn K. et al., Nucleic Acids Res. 2014), we try to structurally character-ize this complex. In our studies we are applying in vivo and in vitro experi-ments including reconstituted DNA replication assays, real time kinetics of protein-DNA complexes, Mass Spectrometry (MS), Atomic Force Microscopy (AFM) and crystallographic analysis.
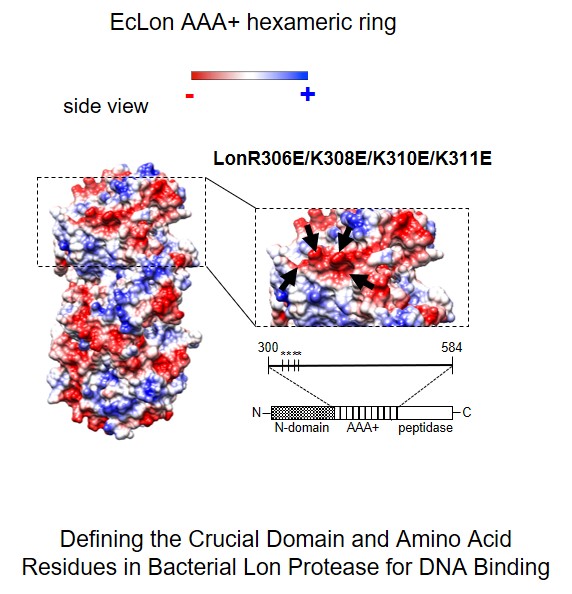
In our research we also challenge the question about the molecular mecha-nism(s) of proteases activity modulation by interaction with nucleic acids. The controlled proteolysis is a long term task as a potential tool in devel-oping anti-microbial therapies. Recently we extended our investigations on the role of proteases in nucleoprotein complex dynamics in bacterial cells. By constructing mutants of Escherichia coli Lon protease, using bioinformatics and phenotypic analysis we demonstrated engagement of positively charged amino acid residues, located on the ATPase domain surface, in interaction with DNA. In vivo tests revealed essentiality of Lon interaction with DNA for Lon activity associated with cell division (Karłowicz A., Węgrzyn K. et. al., JBC 2017).
In the same lane of investigations we tested the influence of proteases on handcuff complex responsible for controlling DNA replication and stable maintenance of plasmids in bacterial cells. We found that bacterial proteas-es, particularly Lon and ClpAP, disrupt handcuff by degrading Rep proteins interacting DNA, thus affecting plasmid stability. Moreover, we demonstrate that Rep monomers are able to dissociate handcuffed plasmid molecules and re-initiate DNA synthesis (Bur K. et. al., NAR 2017). We assume that those mechanisms are important for plasmid maintenance.
We are also developing the line of research considering the mechanisms of replication complex formation in bacterial cells. Our study on the replisome assembly at plasmid RK2 replication origin (Wawrzycka A, et al., Proc Natl Acad Sci USA. 2015) revealed that specific interaction between the plasmid Rep protein and β subunit of DNA Polymerase III holoenzyme (β-clamp) is re-quired for polymerase complex assembly at plasmid origin. Currently conduct-ed experiments are directed towards understanding of the regulatory mecha-nisms involving β-clamp interaction with host and plasmid encoded proteins. We are directing our investigations towards the analysis of the stability of an essential replication proteins and mechanisms involving protease activities for the regulation of both chromosomal and plasmid DNA replication in stress conditions.
Recent and Important Awards:
| 2016 | ELSEVIER-Plasmid prize award for dr. Katarzyna Węgrzyn for the best poster presentation during Plasmid Biology Conference, Cambridge |
| 2015 | Award from The Committee of the Cell Molecular Biology of Polish Academy of Science (PAN) for the best publication in micro-biology published in Polish laboratories |
| 2002 | HHMI (Howard Hughes Medical Institute) Young Investigator |
| 2000 | EMBO YIP (Young Investigator Programme), EMBO Heidelberg |
ONGOING RESEARCH PROJECTS
SONATA NCN (194 172 EUR) started in February 2018
Structural and functional analysis of nucleoprotein complexes of Rep plasmid proteins and the ssDNA of the DUE origin region
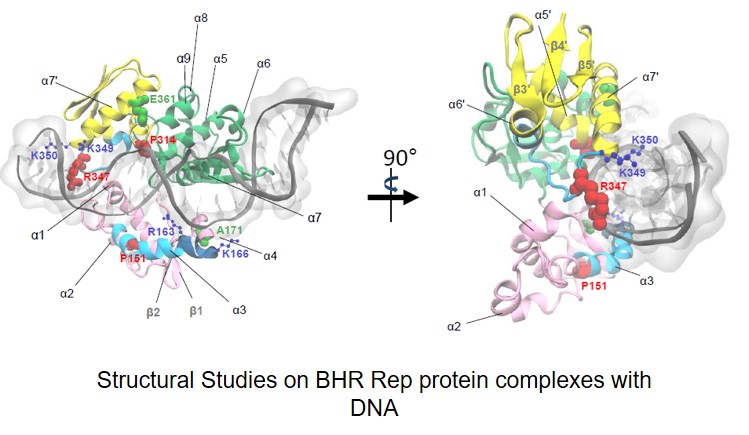
The initiation of DNA replication of extrachromosomal genetic elements, plasmids, is induced by Rep proteins. These proteins contain WH (Winged Helix) domains, which are responsible for dsDNA binding. Our research showed that Reps bind specific region of ssDNA of DUE. However, the structure of the nucleoprotein complexes of Reps and the ssDNA DUE is unknown. The aim of the project is the structural-functional analysis of complexes of plasmid replication initiation proteins, Rep, and the ssDNA DUE. Within this project we plan to determine the domains and the amino acid residues of Rep proteins which are responsible for binding of the ssDNA DUE and to investigate the role of ssDNA DUE binding by Reps. The obtained results should allow to acquire structural data on the formation of nucleoprotein complexes by plasmid replication initiators and to broaden our knowledge concerning the process of DNA replication. Therefore, they will have an impact on the development of the field and might also, in further perspective, have great impact on the development of new antimicrobial strategies.
TEAM FNP (808 874 EUR) started in October 2018
Condition-dependent protease activation for targeted proteolysis in the regulation of DNA replication
The presented project is directed at elucidation how, depending on conditions, proteases are structurally and spatially regulated to degrade essential replication proteins. The research will concentrate on bacterial Lon protease and its phosphate containing cofactors, contributing to the protease activities. We will analyze Lon from bacterial species not being investigated so far in this aspect as well as known and newly identified Lon substrates essential for replication of bacterial DNA. By applying in vitro and in vivo experiments, including real-time, single-molecule and single-cell analysis with a variety of Lon proteins, substrates and cofactors, we expect to describe species-specific, substrate-specific and general effects on Lon, and relate them to growth conditions. The cutting-edge AFM and cryo-EM will be utilize for the description of Lon complexes with cofactors and substrates. Our investigations can provide data on Lon as a new target for new antimicrobial strategies.
RESEARCH TEAM
Laboratory of Molecular Biology
Department of Molecular and Cellular Biology
Intercollegiate Faculty of Biotechnology UG&MUG
Abrahama 58 Street, 80-307 Gdańsk, Poland
Office - Stanisława Urbańska
Tel.:+48 58 523 6421

STAFF LIST
Head: prof. dr hab. Igor Konieczny [link]
tel. +48 58 523 6365 [e-mail]
Post doctoral fellows:
dr hab. Katarzyna Węgrzyn
tel.: +48 58 523 6325 [e-mail]
dr Katarzyna Bury
tel.: +48 58 523 6325 [e-mail]
PhD students:
MSc Weronika Chmura
tel.: +48 58 523 6325 [email]
MSc Zuzanna Hirsz
tel.: +48 58 523 6325 [email]
MSc Ewelina Wysocka
tel.: +48 58 523 6325 [e-mail]
SELECTED PUBLICATIONS
-
Bełdzińska Patrycja, Zakrzewski Marcin, Grzyb Katarzyna, Hauer Amandine, Jamrógiewicz Marzena, Wyrzykowski Dariusz, Bury Katarzyna, Gołuński Grzegorz, Piosik Jacek. Platinum nanoparticles interact with epirubicin in size dependent manner and affect its biological activity. European Journal of Pharmaceutics and Biopharmaceutics 2025, 216: 114866 (doi: 10.1016/j.ejpb.2025.114866)
-
Boguszewska Ewelina, Konieczny Igor. Prevention of RK2 plasmid replication initiation in starved Escherichia coli cells. Plasmid 2025, 134: 102755 (doi: 10.1016/j.plasmid.2025.102755)
-
Bełdzińska Patrycja, Zakrzewski Marcin, Mruk Inez, Bogusławski Marceli, Derewońko Natalia, Bury Katarzyna, Wyrzykowski Dariusz, Gołuński Grzegorz, Rychłowski Michał, Piosik Jacek. Size dependent impact of platinum nanoparticles on doxorubicin activity, European Journal of Pharmaceutical Sciences, 2025, 3:107094 (doi:10.1016/j.ejps.2025.107094)
-
Bełdzińska Patrycja, Galikowska-Bogut Barbara, Zakrzewski Marcin, Bury Katarzyna, Jamrógiewicz Marzena, Wyrzykowski Dariusz, Gołuński Grzegorz, Sądej Rafał, Piosik Jacek. Platinum as both a drug and its modulator - Do platinum nanoparticles influence cisplatin activity? Chemico-Biological Interactions 2025, 407: 111365 (doi: 10.1016/j.cbi.2024.111365)
-
Orlikowska Marta, Wyciszkiewicz Aleksandra, Węgrzyn Katarzyna, Mehringer Johannes, de Souza Paiva Daisylea, Jurczak Przemysław. Methods for monitoring protein-membrane binding. Comparison based on the interactions between amyloidogenic protein human cystatin C and phospholipid liposomes. International Journal of Biological Macromolecules 2024, 278(3): 134889 (doi: 10.1016/j.ijbiomac.2024.134889)
-
Gołuński Grzegorz, Konkel Kinga, Galikowska-Bogut Barbara, Bełdzińska Patrycja, Bury Katarzyna, Zakrzewski Marcin, Butowska Kamila, Sądej Rafał, Piosik Jacek. Infuence of silver nanoparticles’ size on their direct interactions with doxorubicin and its biological effects. Scientific Reports 2024, 14: 18544 (doi: 10.1038/s41598-024-69724-6)
-
Ridgway Harry, Moore Graham J., Gadanec Laura Kate, Zulli Anthony, Apostolopoulos Vasso, Hoffmann Weronika, Węgrzyn Katarzyna, Vassilaki Niki, Mpekoulis George, Zouridakis Marios, Giastas Petros, Vidali Veroniki P., Kelaidonis Konstantinos, Matsoukas Minos-Timotheos, Dimitriou Marios, Mavromoustakos Thomas, Tsiodras Sotirios, Gorgoulis Vasilis G., Karakasiliotis Ioannis, Chasapis Christos T., Matsoukas John M. Novel benzimidazole angiotensin receptor blockers with anti-SARS-CoV-2 activity equipotent to that of nirmatrelvir: computational and enzymatic studies. Expert Opinion on Therapeutic Targets 2024, 28(5): 437-459 (doi: 10.1080/14728222.2024.2362675)
-
Kocikowski Mikołaj, Dziubek Katarzyna, Węgrzyn Katarzyna, Hrabal Václav, Zavadil-Kokas Filip, Vojtesek Borivoj, Alfaro Javier Antonio, Hupp Theodore, Parys Maciej. Comparative characterization of two monoclonal antibodies targeting canine PD-1. Frontiers in Immunology 2024, 15: 1382576 (doi: 10.3389/fimmu.2024.1382576)
-
Bojko Magdalena, Węgrzyn Katarzyna, Sikorska Emilia, Ciura Piotr, Battin Claire, Steinberger Peter, Magiera-Mularz Katarzyna, Dubin Grzegorz, Kulesza Adam, Sieradzan Adam, Spodzieja Marta, Rodziewicz-Motowidło Sylwia. Peptide-based inhibitors targeting the PD-1/PD-L1 axis: potential immunotherapeutics for cancer. Translational Oncology 2024, 42: 101892 (doi: 10.1016/j.tranon.2024.101892)
- Węgrzyn Katarzyna, Konieczny Igor. Toward an understanding of the DNA replication initiation in bacteria. Frontiers in Microbiology 2024, 14: 1328842 (doi: 10.3389/fmicb.2023.1328842)
-
Borowicz Marcin, Krzyżanowska Dorota, Narajczyk Magdalena., Sobolweska Marta, Rajewska Magdalena, Czaplewska Paulina, Węgrzyn Katarzyna, Czajkowski Robert. Soft rot pathogen Dickeya dadantii 3937 produces tailocins resembling the tails of Peduovirus P2. Frontiers in Microbiology 2023, 14 (doi: 10.3389/fmicb.2023.1307349)
-
Brankiewicz Wioletta, Kalathiya Umesh, Padariya Monikaben, Węgrzyn Katarzyna, Prusinowski Maciej, Żebrowska Joanna, Żylicz-Stachula Agnieszka, Skowron Piotr, Drab Marek, Szajewski Mariusz, Ciesielski Maciej, Gawrońska Małgorzata, Kallingal Anoop, Makowski Mariusz, Bagiński Maciej. Modified peptide molecules as potential modulators of shelterin protein functions; TRF1. Chemistry-A European Journal 2023, 29(55): e202300970 (doi: 10.1002/chem.202300970)
-
Węgrzyn Katarzyna, Oliwa Monika, Nowacka Marzena, Zabrocka Elżbieta, Bury Katarzyna, Purzycki Piotr, Czaplewska Paulina, Pipka Justyna, Giraldo Rafael, Konieczny Igor. Rep protein accommodates together dsDNA and ssDNA which enables a loop-back mechanism to plasmid DNA replication initiation. Nucleic Acids Research 2023, 51(19): 10551-10567 (doi: 10.1093/nar/gkad740)
-
Kuncewicz Katarzyna, Bojko Magdalena, Battin Claire, Karczyńska Agnieszka, Sieradzan Adam, Sikorska Emilia, Węgrzyn Katarzyna, Wojciechowicz Karolina, Wardowska Anna, Steinberger Peter, Rodziewicz-Motowidło Sylwia, Spodzieja Marta. BTLA-derived peptides as inhibitors of BTLA/HVEM complex formation – design, synthesis and biological evaluation. Biomedicine & Pharmacotherapy 2023, 165: 115161 (doi: 10.1016/j.biopha.2023.115161)
-
Kelaidonis Konstantinos, Ligielli Irene, Letsios Spiros, Vidali Veroniki, Mavromoustakos Thomas, Vassilaki Niki, Moore Graham, Hoffmann Weronika, Węgrzyn Katarzyna, Ridgway Harry, Chasapis Christos, Matsoukas John. Computational and enzymatic studies of sartans in SARS-CoV-2 spike RBD-ACE2 binding: the role of tetrazole and perspectives as antihypertensive and COVID-19 therapeutics. International Journal of Molecular Sciences 2023, 24(9): 8454 (doi: 10.3390/ijms24098454)
-
Machón Cristina, Ruiz-Masó José A., Amodio Juliana, Boer D. Roeland, Bordanaba-Ruiseco Lorena, Bury Katarzyna, Konieczny Igor, del Solar Gloria, Coll Miquel. Structures of pMV158 replication initiator RepB with and without DNA reveal a flexible dual-function protein. Nucleic Acids Research 2023, 51(3): 1458-1472 (doi: 10.1093/nar/gkac1271)
-
Rybicka Magda, Czaplewska Paulina, Rzymowska Jolanta, Weronika Sofińska-Chmiel, Wójcik-Mieszawska Sylwia, Lewtak Kinga, Węgrzyn Katarzyna, Jurczak Przemysław, Szpiech Agata, Nowak Jakub, Musiał Natalia, Fiołka Marta J.. Novel Venetin-1 nanoparticle from earthworm coelomic fluid as a promising agent for the treatment of non-small cell lung cancer. Scientific Reports 2022, 12(1): 18497 (doi: 10.1038/s41598-022-21665-8)
-
Bojko Magdalena, Węgrzyn Katarzyna, Sikorska Emilia, Kocikowski Mikołaj, Parys Maciej, Battin Claire, Steinberger Peter, Kogut Małgorzata, Winnicki Michał, Sieradzan Adam, Spodzieja Marta, Rodziewicz-Motowidło Sylwia. Design, synthesis and biological evaluation of PD-1 derived peptides as inhibitors of PD-1/PD-L1 complex formation for cancer therapy. Bioorganic Chemistry 2022, 128: 106047 (doi: 10.1016/j.bioorg.2022.106047)
-
Kadziński Leszek, Łyżeń Robert, Bury Katarzyna, Banecki Bogdan. Modeling and optimization of β-galactosidase entrapping in polydimethylsiloxane-modified silica composites. International Journal of Molecular Sciences 2022, 23(10): 5395 (doi: 10.3390/ijms23105395)
-
Kuncewicz Katarzyna, Battin Claire, Węgrzyn Katarzyna, Sieradzan Adam, Wardowska Anna, Sikorska Emilia, Giedrojć Irma, Smardz Pamela, Pikuła Michał, Steinberger Peter, Rodziewicz-Motowidło Sylwia, Spodzieja Marta. Targeting the HVEM protein using a fragment of glycoprotein D to inhibit formation of the BTLA/HVEM complex. Bioorganic Chemistry 2022, 122: 105748 (doi: 10.1016/j.bioorg.2022.105748)
-
Węgrzyn Katarzyna, Konieczny Igor. Archaeal Orc1 protein interacts with T-rich single-stranded DNA. BMC Research Notes 2021, 14(1): 275 (doi: 10.1186/s13104-021-05690-w)
-
Kołodziej-Sobocińska Marta, Trojanowski Damian, Bury Katarzyna, Hołówka Joanna, Matysik Weronika, Kąkolewska Hanna, Feddersen Helge, Giacomelli Giacomo, Konieczny Igor, Bramkamp Marc, Zakrzewska-Czerwińska Jolanta. Lsr2, a nucleoidassociated protein influencing mycobacterial cell cycle. Scientific Reports 2021, 11(1): 1-12 (doi: 10.1038/s41598-021-82295-0)
-
Kołodziej-Sobocińska Marta, Łebkowski Tomasz, Płociński Przemysław, Hołówka Joanna, Paściak Mariola, Wojtaś Bartosz, Bury Katarzyna, Konieczny Igor, Dziadek Jarosław, Zakrzewska-Czerwińska Jolanta. Lsr2 and its novel paralogue mediate the adjustment of Mycobacterium smegmatis to unfavorable environmental conditions. mSphere 2021, 6(3): 1-18 (doi: 10.1128/msphere.00290-21)
-
Węgrzyn Katarzyna, Zabrocka Elżbieta, Bury Katarzyna, Tomiczek Bartłomiej, Wieczór Miłosz, Czub Jacek, Uciechowska-Kaczmarzyk Urszula, Moreno-del Alamo Maria, Walkow Urszula, Grochowina Igor, Dutkiewicz Rafał, Bujnicki Janusz M., Giraldo Rafael, Konieczny Igor. Defining a novel domain that provides an essential contribution to site-specific interaction of Rep protein with DNA. Nucleic Acids Research 2021, 49(6): 3394-3408 (doi: 10.1093/nar/gkab113)
-
Romanowska Anita, Węgrzyn Katarzyna, Bury Katarzyna, Sikorska Emilia, Gnatek Aleksandra, Piwkowska Agnieszka, Konieczny Igor, Lesner Adam, Wysocka Magdalena. Novel cell permeable polymers of N-substituted L-2,3-diaminopropionic acid (DAPEGs) and cellular consequences of their interactions with nucleic acids. International Journal Of Molecular Sciences 2021, 22(5): 2571 (doi: 10.3390/ijms22052571)
-
Ropelewska Małgorzata, Gross Marta H., Konieczny Igor. DNA and Polyphosphate in Directed Proteolysis for DNA Replication Control. Frontiers in Microbiology 2020, 11: 585717 (doi: 10.3389/fmicb.2020.585717)
-
Gross H. Marta, Konieczny Igor. Polyphosphate induces the proteolysis of ADP-bound fraction of initiator to inhibit DNA replication initiation upon stress in Escherichia coli. Nucleic Acids Research 2020, 48(10): 5457-5466 (doi: 10.1093/nar/gkaa217)
-
Mróz Dagmara, Wyszkowski Hubert, Szablewski Tomasz, Zawieracz Katarzyna, Dutkiewicz Rafał, Bury Katarzyna, Wortmann Saskia B., Wevers Ron A., Ziętkiewicz Szymon. CLPB (caseinolytic peptidase B homolog), the first mitochondrial protein refoldase associated with human disease. Biochimica et Biophysica Acta (BBA) - General Subjects 2020, 1864 (4): 129512 (doi: 10.1016/j.bbagen.2020.129512)
-
Spodzieja Marta, Kuncewicz Katarzyna, Sieradzan Adam, Karczyńska Agnieszka, Iwaszkiewicz Justyna, Cesson Valérie, Węgrzyn Katarzyna, Zhukov Igor, Maszota-Zieleniak Martyna, Michielin Olivier, Speiser Daniel E., Zoete Vincent, Derré Laurent, Rodziewicz-Motowidło Sylwia. Disulfide-Linked Peptides for Blocking BTLA/HVEM Binding. International Journal of Molecular Sciences 2020, 21(2): 636 (doi: 10.3390/ijms21020636)
-
Colizzi Vittorio, Mezzana Daniele, Ovseiko Pavel, V., Caiati Giovanni, Colonnello Claudia, Declich Andrea, Buchan Alastair M., Edmunds Laurel, Buzan Elena, Zerbini Luiz, Djilianov Dimitar, Schmidt Evanthia Kalpazidou, Bielawski Krzysztof, Elster Doris, Solvato Maria, Alcantara Luiz C., Jr., Minutolo Antonella, Potesta Marina, Bachiddu Elena, Milano Maria J., Henderson Lorna R., Kiparoglou Vasiliki, Friesen Phoebe, Sheehan Mark, Moyankova Daniela, Rusanov Krasimir, Wium Martha, Raszczyk Izabela, Konieczny Igor, Gwizdala Jerzy P., Sledzik Karol, Barendziak Tanja, Birkholz Julia, Mueller Nicklas, Warrelmann Juergen, Meyer Ute, Filser Juliane, Barreto Fernando Khouri, Montesano Carla. Structural Transformation to Attain Responsible BIOSciences (STARBIOS2): Protocol for a Horizon 2020 Funded European Multicenter Project to Promote Responsible Research and Innovation. JMIR Research Protocols 2019, 8 (3): e11745 (doi: 10.2196/11745)
-
Dubiel Andrzej, Węgrzyn Katarzyna, Kupiński Adam P., Konieczny Igor. ClpAP protease is a universal factor that activates the parDE toxin-antitoxin system from a broad host range RK2 plasmid. Scientific Reports 2018, 8: 15287 (doi: 10.1038/s41598-018-33726-y)
-
Bury Katarzyna, Węgrzyn Katarzyna, Konieczny Igor. Handcuffing reversal is facilitated by proteases and replication initiator monomers. Nucleic Acids Research 2017, 45(7): 3953-3966 (doi: 10.1093/nar/gkx166)
-
Karłowicz Anna, Węgrzyn Katarzyna, Gross Marta, Kaczyńska Dagmara, Ropelewska Małgorzata, Siemiątkowska Małgorzata, Bujnicki Janusz, Konieczny Igor. Defining the Crucial Domain and Amino Acid Residues in Bacterial Lon Protease for DNA Binding and Processing of DNA-interacting Substrates Journal of Biological Chemistry 2017, 292: 7507-7518 (doi:10.1074/jbc.M116.766709)
-
Karłowicz Anna, Węgrzyn Katarzyna, Dubiel Andrzej, Ropelewska Małgorzata, Konieczny Igor. Proteolysis in plasmid DNA stable maintenance in bacterial cells. Plasmid 2016, 86: 7-13 (doi: 10.1016/j.plasmid.2016.05.002)
-
Węgrzyn Katarzyna E., Gross Marta, Uciechowska Urszula, Konieczny Igor. Replisome Assembly at Bacterial Chromosomes and Iteron Plasmids. Frontiers in Molecular Biosciences 2016, 3: 39 (doi: 10.3389/fmolb.2016.00039)
-
Yano Hirokazu, Wegrzyn Katarzyna, Loftie-Eaton Wesley, Johnson Jenny Deckert Gail E., Rogers Linda M., Konieczny Igor, Top Eva M. Evolved plasmid-host interactions reduce plasmid interference cost. Molecular Microbiology, 2016, 101(5): 743-756 (doi: 10.1111/mmi.13407)
-
Koper Tomasz, Polit Agnieszka, Sobiecka-Szkatula Anna, Węgrzyn Katarzyna, Scire Andrea, Figaj Donata, Kadziński Leszek, Zarzecka Urszula, Żurawa-Janicka Dorota, Banecki Bogdan, Lesner Adam, Tanfani Fabio, Lipińska Barbara, Skórko-Glonek Joanna. Analysis of the Link between the Redox State and Enzymatic Activity of the HtrA (DegP) Protein from Escherichia coli. PLoS ONE 2015, art. no e0117413 (1-24) (doi: 10.1371/journal.pone.0117413).
-
Wawrzycka Aleksandra, Gross Marta, Wasążnik Anna, Konieczny Igor. Plasmid replication initiator interactions with origin 13-mers and polymerase subunits contribute to strand-specific replisome assembly. Proceedings of the National Academy of Sciences of the United States of America 2015, 112(31): E4188-E4196 (doi: 10.1073/pnas.1504926112).
-
Lindgren Cecilia, Andersson Ida E., Berg Lotta, Dobritzsch Doreen, GeChangrong, Haag Sabrina, Uciechowska Urszula, Holmdahl Rikard, Kihlberg Jan, Linusson Anna. Hydroxyethylene isosteres introduced in type II collagen fragments substantially alter the structure and dynamics of class II MHC Aq /glycopeptide complexes. Organic & Biomolecular Chemistry 2015, 13(22):
6203-6216 (doi: 10.1039/c5ob00395d) -
Konieczny Igor, Bury Katarzyna, Wawrzycka Aleksandra, Węgrzyn Katarzyna. Iteron plasmids. Microbiology Spectrum 2014, 2(6): art. no PLAS-0026-2014 (1-16) (doi: 10.1128/microbiolspec.PLAS-0026-2014).
-
Osbourne Devon O., Soo Valerie W. C., Konieczny Igor, Wooda Thomas K. Polyphosphate, cyclic AMP, guanosine tetraphosphate, and c-di-GMP reduce in vitro Lon activity. Bioengineered 2014, 5(4): 264-268 (doi: 10.4161/bioe.29261).
-
Węgrzyn Katarzyna, Fuentes-Perez Maria Eugenia, Bury Katarzyna, Rajewska Magdalena, Moreno-Herrero Fernando, Konieczny Igor. Sequence-specific interactions of Rep proteins with ssDNA in the AT-rich region of the plasmid replication origin. Nucleic Acids Research 2014, 42(12): 7807-7818 (doi: 10.1093/nar/gku453).
-
Zabrocka Elżbieta, Węgrzyn Katarzyna, Konieczny Igor. Two replication initiators - one mechanism for replication origin opening? Plasmid 2014, 76: 72-78 (doi: 10.1016/j.plasmid.2014.10.003).
-
Diago-Navarro Elizabeth, Hernandez-Arriaga Ana Maria, Kubik Sławomir, Konieczny Igor, Diaz-Orejas Ramon. Cleavage of the antitoxin of the parD toxin-antitoxin system is determined by the ClpAP protease and is modulated by the relative ratio of the toxin and the antitoxin. Plasmid 2013, 70(1): 78-85
-
Węgrzyn Katarzyna, Witosińska Monika, Schweiger Paweł, Bury Katarzyna, Jenal Urs, Konieczny Igor. RK2 plasmid dynamics in Caulobacter crescentus cells - two modes of DNA replication initiation. Microbiology-SGM 2013, 159(Pt 6): 1010-1022
-
Kubik Sławomir, Węgrzyn Katarzyna, Pierechod Marcin, Konieczny Igor. Opposing effects of DNA on proteolysis of a replication initiator. Nucleic Acids Research 2012, 40(3): 1148-1159
-
Rajewska Magdalena, Węgrzyn Katarzyna, Konieczny Igor. AT-rich region and repeated sequences - the essential elements of replication origins of bacterial replicons. FEMS Microbiology Reviews 2012, 36(2): 408-434
-
Szambowska Anna, Pierechod Marcin, Węgrzyn Grzegorz, Glinkowska Monika. Coupling of transcription and replication machineries in λ DNA replication initiation: evidence for direct interaction of Escherichia coli RNA polymerase and the λO protein. Nucleic Acids Research 2011, 39(1): 168-177
-
Kołatka Katarzyna, Kubik Sławomir, Rajewska Magdalena, Konieczny Igor. Replication and partitioning of the broad-host-range plasmid RK2. Plasmid 2010, 64(3): 119-134
-
Pierechod Marcin, Nowak Agnieszka, Saari Anna, Purta E., Bujnicki Janusz M., Konieczny I. Conformation of a plasmid replication initiator protein affects its proteolysis by ClpXP system. Protein Science 2009, 18(3): 637-649




